AI driven antibody sequence generation
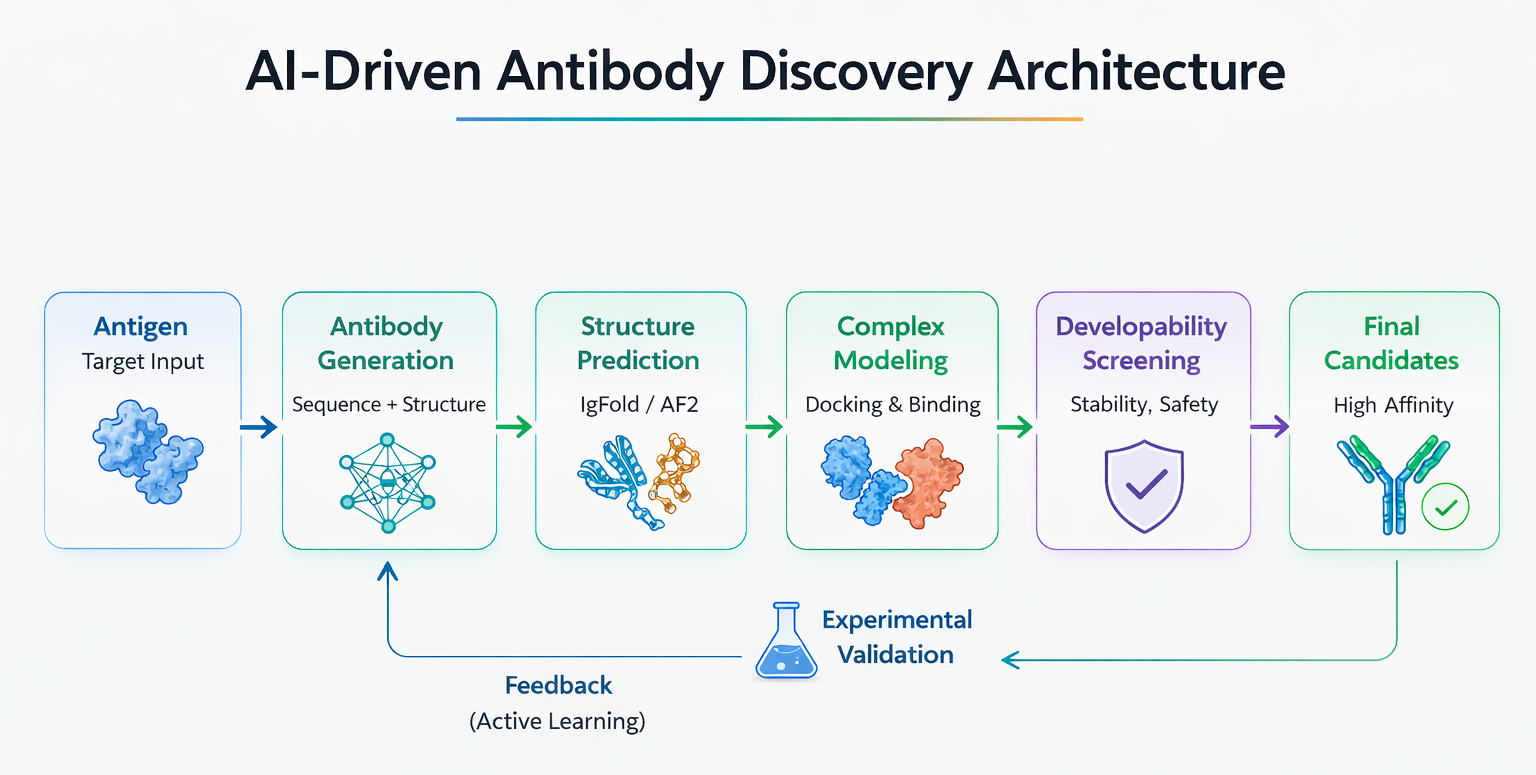
Developed an AI driven pipeline for antibody sequence generation, leveraging deep learning and generative models trained on large scale OAS & SAbDab antibody datasets. Integrated with sequence optimization and structural validation tools to design high affinity, developable antibodies with therapeutic potential.
AntiFold | AbLang | diffAb | BioPhi | LLM fine tuning
Knowledge Graph
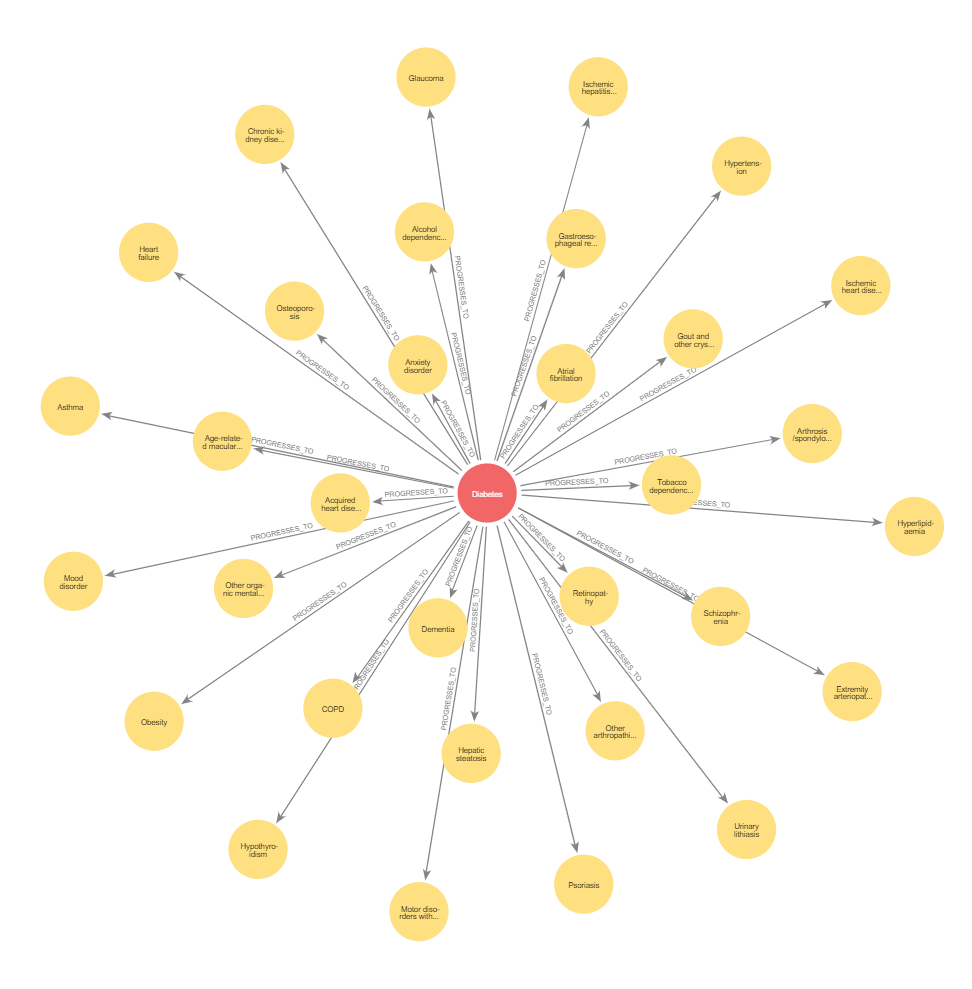
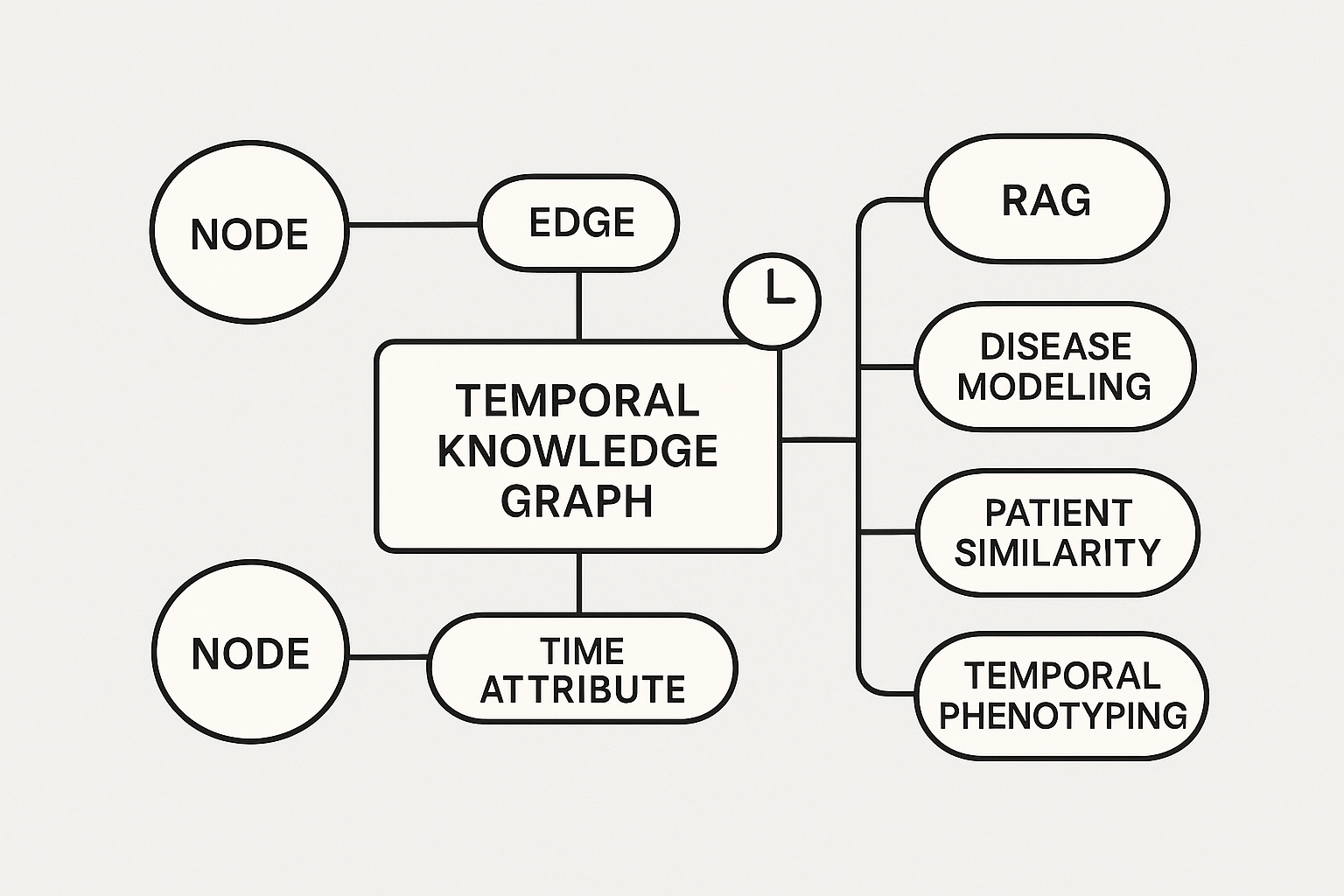
Built a KG with open source biological/Real World/Clinical data, harmonized with controlled vocabularies for each entity. Application included drug repurposing, target identification, safety assessment for toxicity and organ wise stratification, reducing months of work to weeks
Neo4j | Link prediction | Node classification | Community Detection
Automated QSPR model building for property prediction
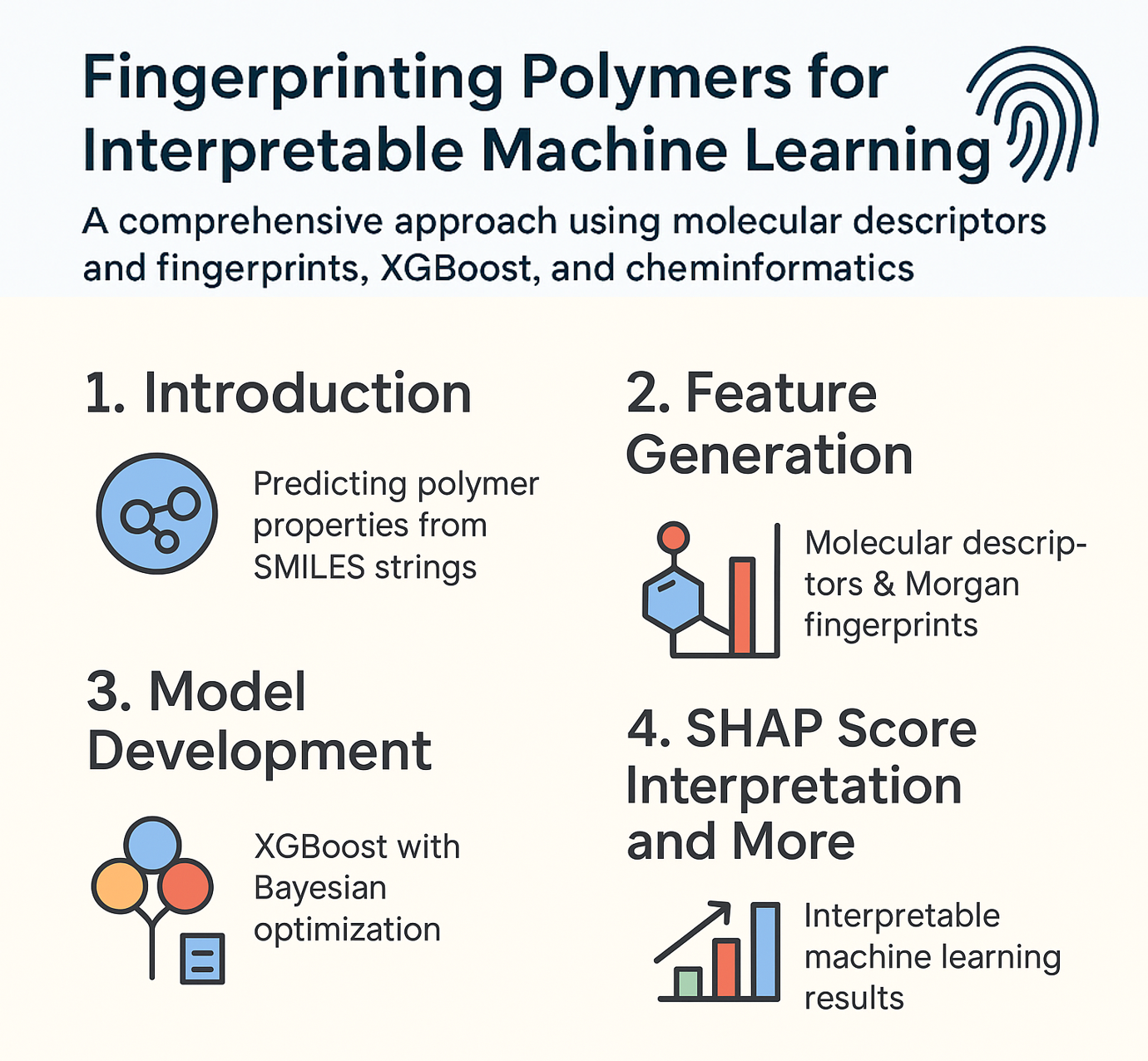
Developed an automated ML pipeline to build Quantitative Structure-Property Relationship (QSPR) model for drug property prediction, that helped reducing dependency on data scientist for model building and increased capabilities across departments.
RDKit | Morgan fingerprint | Optuna - Bayesian Optimization
Machine Learning driven biomarker identification
Developed a computational pipeline for automated biomarker identification using TCGA database, by training classical ML models using RNA seq data, to identify diagnostic and prognostic signatures in BRCA, LIHC, LUAD and PRAD. Enabling faster patient stratification for precision medicine applications.
DEG analysis | WCGNA | Applied ML | SHAP | Kaplan Meier
Network analysis - a systems approach for target identification and prioritization
1. High throughput drug combination (synergy) assessment using network proximity analysis
2. Network vulnerability analysis for target prioritization
Network proximity | Network attributes | STRING
GWAS - WGS
1. GenomeIndia cohort: Performed quality control and variant association analysis followed by cross population studies.
2. AllOfUs cohort: Developed a Cromwell (WDL) pipeline to scale up association studies across 3000+ traits using EHRs to identify statistically significant biomarkers. Association statistics for the study included dosage sensitivity and gene burden test.
Association statistics | Dosage sensitivity | Burden test
In silico KnockOut/perturbation
Developed a high throughput Boolean model simulation pipeline for in silico gene knockout/perturbation experiments, using RNA seq data to initialize the system states supporting data-driven therapeutics which enhances precision in target prioritization.
Attractors | Trap spaces | Logical gates | KEGG
Structure based druggability prediction
Developed a structure based druggability prediction pipeline leveraging parallel processing to accelerate searches across a database of known binding pockets, enabling rapid identification of similar sites to assess target protein druggability.
Graph Analytics | Structural bioinformatics | Multiprocessing
Data driven spectral deconvolution and compound detection
Developed an automated compound identification pipeline leveraging spectral data, matched against a processed in-house reference compound database.
Spectral data processing | Automation